Load data, estimate and compare models
load_estimate_compare_models.RmdImporting a .mom file as a gnssts
object
Let us first load the gmwmx package.
library(gmwmx)Consider that you want to estimate a model on data saved in a
.mom file located at a specific file_path on
your computer, where file_path is the path where is located
the .mom file (for example
file_path = "/home/name_of_the_user/Documents/data.mom")
For example, the corresponding .mom file could have a
similar looking:
# sampling period 1.000000
# offset 55285.000000
# offset 58287.770833
52759.5 -0.01165
52760.5 -0.01102
52761.5 -0.01147
...You can import the .mom file as a with the function
read.gnssts() as such:
data_dobs = read.gnssts(filename = file_path)Objects created or imported with create.gnss() or
read.gnssts() are of class gnssts.
class(data_dobs)## [1] "gnssts"By inspecting the structure of a gnssts object, we
observe that gnssts objects specify the time vector, the
observation vector, the sampling period and the times at which there are
location shifts (jumps).
str(data_dobs)## List of 4
## $ t : num [1:5559] 52760 52760 52762 52764 52766 ...
## $ y : num [1:5559] -0.0117 -0.011 -0.0115 -0.0131 -0.0106 ...
## $ sampling_period: num 1
## $ jumps : num [1:2] 55285 58288
## - attr(*, "class")= chr "gnssts"We can represent the signal as such:
plot(data_dobs$t, data_dobs$y, type="l")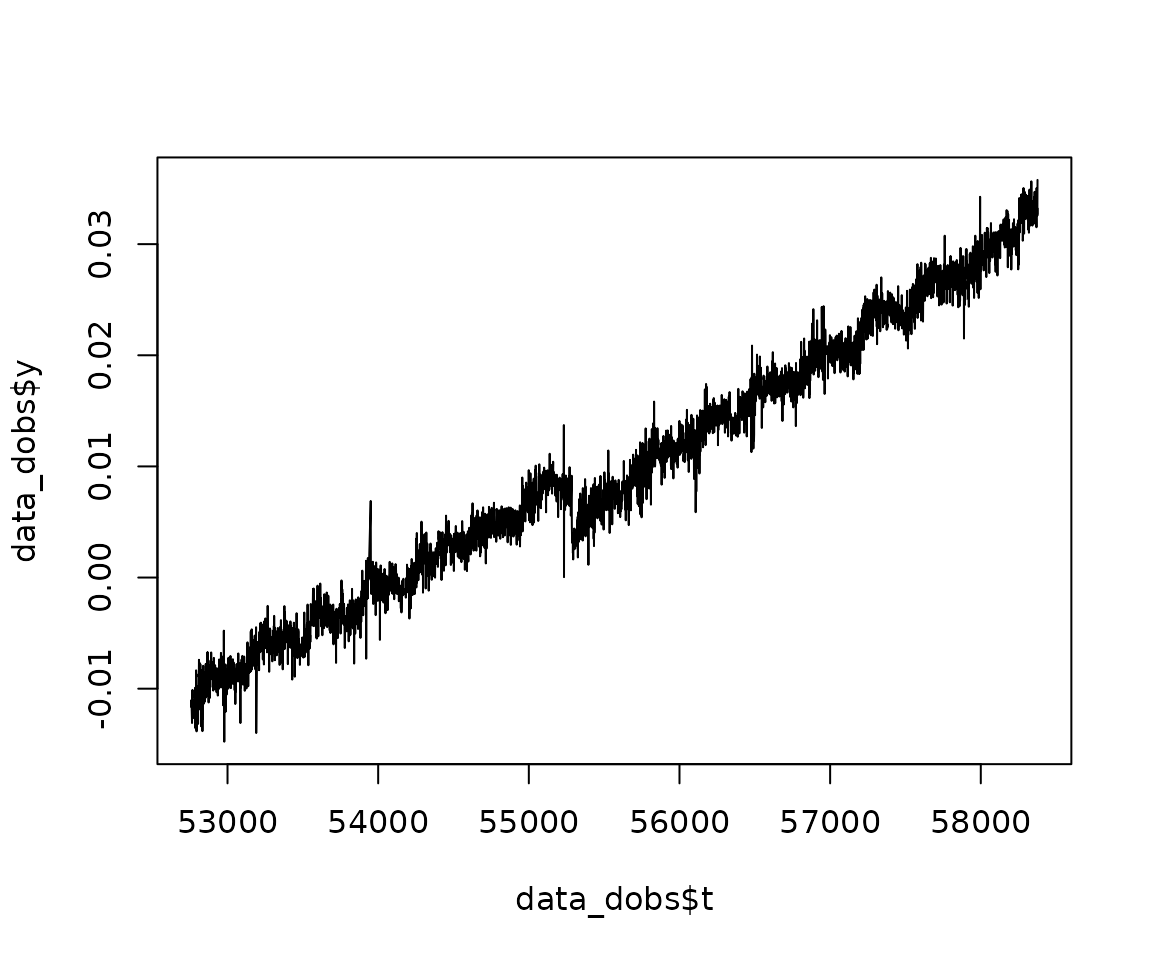
Estimate a model
Supported models
The gmwmx package allows to estimate linear model with
correlated residuals that are described by a functional model and a
stochastic noise model.
Functional model
More precisely, for the functional model, we consider a linear model which can be expressed as:
$$\begin{equation} \mathbf{Y} = \mathbf{A} {{\bf x}}_0 + \boldsymbol{\varepsilon}, \end{equation}$$
where $\mathbf{Y} \in {\rm I\!R}^n$ denotes the response variable of interest (i.e., vector of GNSS observations), $\mathbf{A} \in {\rm I\!R}^{n \times p}$ a fixed design matrix, ${{\bf x}}_0 \in \mathcal{X} \subset {\rm I\!R}^p$ a vector of unknown constants and $\boldsymbol{\varepsilon} \in {\rm I\!R}^n$ a vector of (zero mean) residuals.
The gmwmx package allows to estimate functional models
for which the
-th
component of the vector $\mathbf{A} {{\bf
x}}_0$ can be described as follows:
$$\begin{equation} \mathbb{E}[\mathbf{Y}_i] = \mathbf{A}_i^T {{\bf x}}_0 = a+b\left(t_{i}-t_{0}\right)+\sum_{h=1}^{2}\left[c_{h} \sin \left(2 \pi f_{h} t_{i}\right)+d_{h} \cos \left(2 \pi f_{h} t_{i}\right)\right] + \sum_{k=1}^{n_{g}} g_{k} H\left(t_{i}-t_{k}\right), \end{equation}$$
where is the initial position at the reference epoch , is the velocity parameter, and are the periodic motion parameters ( and represent the annual and semi-annual seasonal terms, respectively). The offset terms models earthquakes, equipment changes or human intervention in which is the magnitude of the change at epochs , is the total number of offsets, and is the Heaviside step function. Note that the estimates of the parameters of the functional model are provided in unit/day.
Stochastic model
Regarding the stochastic model, we assume that is a strictly (intrinsically) stationary process and that
where denotes some probability distribution in ${\rm I\!R}^n$ with mean ${\bf 0}$ and covariance .
We assume that and that it depends on the unknown parameter vector $\boldsymbol{\gamma}_0 \in \boldsymbol{\Gamma} \subset {\rm I\!R}^q$. This parameter vector specifies the covariance of the observations and is often referred to as the stochastic parameters.
Hence, we let $\boldsymbol{\theta}_0 = \left[\boldsymbol{{\bf x}}_0^{\rm T} \;\; \boldsymbol{\gamma}_0^{\rm T}\right]^{\rm T} \in \boldsymbol{\Theta} = \mathcal{X} \times \boldsymbol{\Gamma} \subset {\rm I\!R}^{p + k}$ denote the unknown parameter vector of the model described above.
The gmwmx allows to estimate parameters of a specified
functional model as well as parameters of a stochastic model
(i.e. )
defined by a combinations of
- White noise (
wn) - Matérn process (
matern) - Fractional Gaussian noise (
fgn) and - Power Law process (
powerlaw).
Note that only the gmwmx current version accepts only
one process of each kind.
Estimate a model with the GMWMX
You can estimate a model using the GMWMX estimator with the function
estimate_gmwmx().
The stochastic model considered is specified by a string provided to
the argument model_string which is a combination of the
strings wn, powerlaw, matern and
fgn separated by the character +.
You specify the initialization values for solving the optimization
problem at the GMWM estimation step that estimate the stochastic model
by providing a numeric vector of the correct length (the total number of
parameters of the stochastic model specified in
model_string) to the argument theta_0.
You can compute confidence intervals for estimated functional
parameters of an estimated model by setting the argument ci
to TRUE.
Let us consider a single sinusoidal signal with the jumps specified
in the gnssts object and a combination of a White noise and
a Power Law process for the stochastic model.
fit_dobs_wn_plp_gmwmx = estimate_gmwmx(x = data_dobs, theta_0 = c(0.1, 0.1, 0.1),
model_string = "wn+powerlaw",
n_seasonal = 1, ci = T)Estimated models are of class gnsstsmodel
class(fit_dobs_wn_plp_gmwmx)## [1] "gnsstsmodel"We can print the estimated model or extract estimated parameters (functional and stochastic) as such:
print(fit_dobs_wn_plp_gmwmx)## GNSS time series model
##
## * Model: wn + powerlaw
##
## * Functional parameters:
## bias : +0.012013 +/- 0.0200366449
## trend : +0.000008 +/- 0.0000001991
## A*cos(U) : -0.000655 +/- 0.0000754528
## A*sin(U) : +0.000016 +/- 0.0000762463
## jump : -0.003758 +/- 0.0005738092
## jump : +0.001568 +/- 0.0006045750
##
## * Stochastic parameters:
## wn_sigma2 : +0.00000079
## powerlaw_sigma2 : +0.00000025
## powerlaw_d : +0.49990000
##
## * Estimation time: 1.66 s
fit_dobs_wn_plp_gmwmx$beta_hat## bias trend A*cos(U) A*sin(U) jump
## 1.201297e-02 8.351512e-06 -6.553985e-04 1.578605e-05 -3.758074e-03
## jump
## 1.567881e-03
fit_dobs_wn_plp_gmwmx$theta_hat## wn_sigma2 powerlaw_sigma2 powerlaw_d
## 7.916985e-07 2.525867e-07 4.999000e-01We can also plot graphically the estimated functional model on the
time series and the Wavelet variance of residuals by calling the
plot.gnsstsmodel method on a gnsstsmodel
object.
plot(fit_dobs_wn_plp_gmwmx)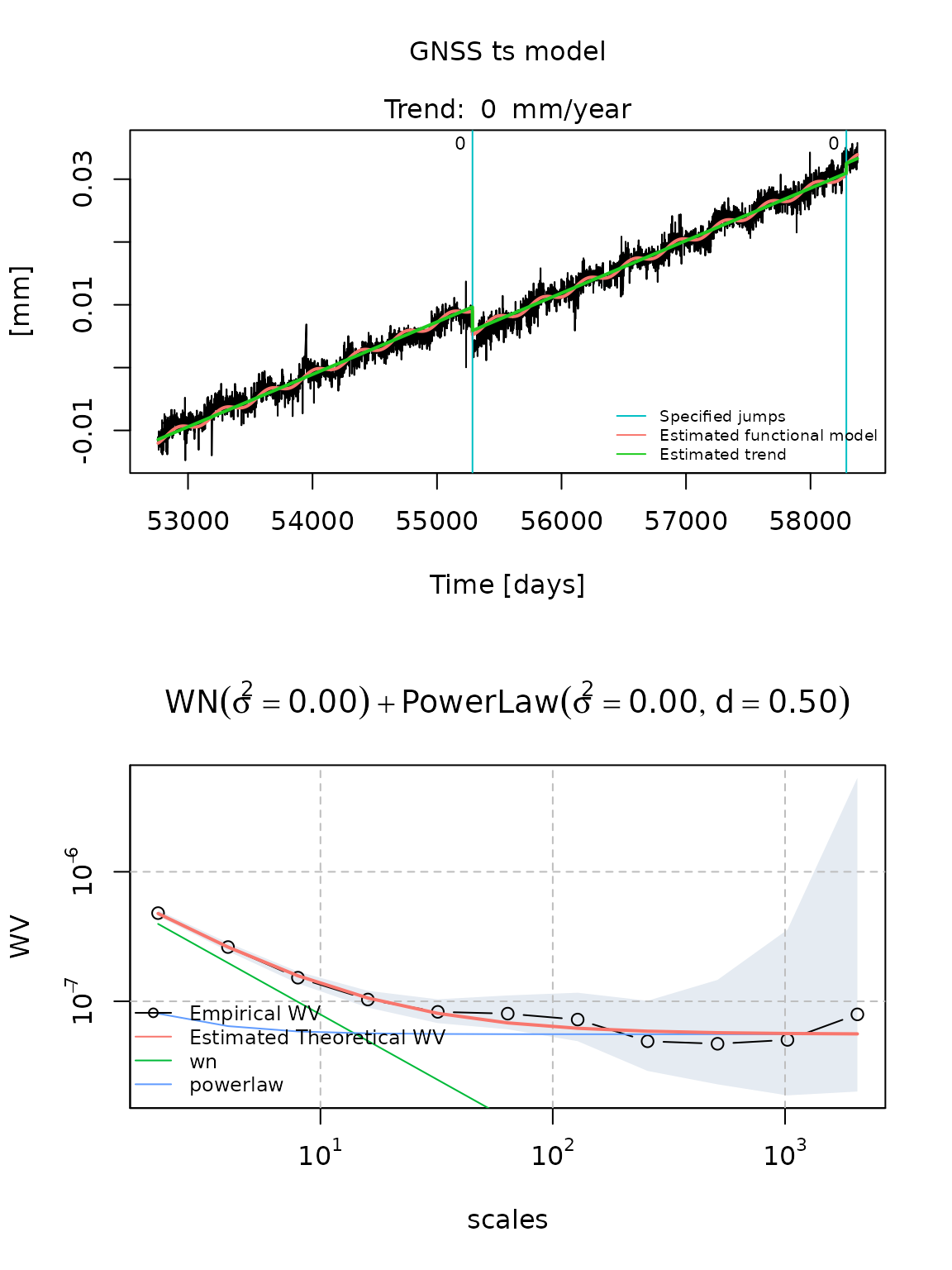
We can specify the number of iterations of the GMWMX to compute
respectively the GMWMX-1 and GMWMX-2 or other iteration of the GMWMX
with the argument k_iter. For example we can compute the
GMWMX-2 as such:
fit_dobs_wn_plp_gmwmx_2 = estimate_gmwmx(x = data_dobs,
theta_0 = c(0.1, 0.1, 0.1),
model_string = "wn+powerlaw",
n_seasonal = 1,
k_iter = 2)Estimate a model with the MLE implemented in Hector
Assuming that you have Hector available on the PATH, an
estimation of the model can the be performed using the Maximum
Likelihood Estimation (MLE) method implemented in Hector as such:
fit_dobs_wn_plp_mle = estimate_hector(x = data_dobs,
model_string = "wn+powerlaw",
n_seasonal = 1)Similarly we can plot and extract the model parameters of the estimated model:
plot(fit_dobs_wn_plp_mle)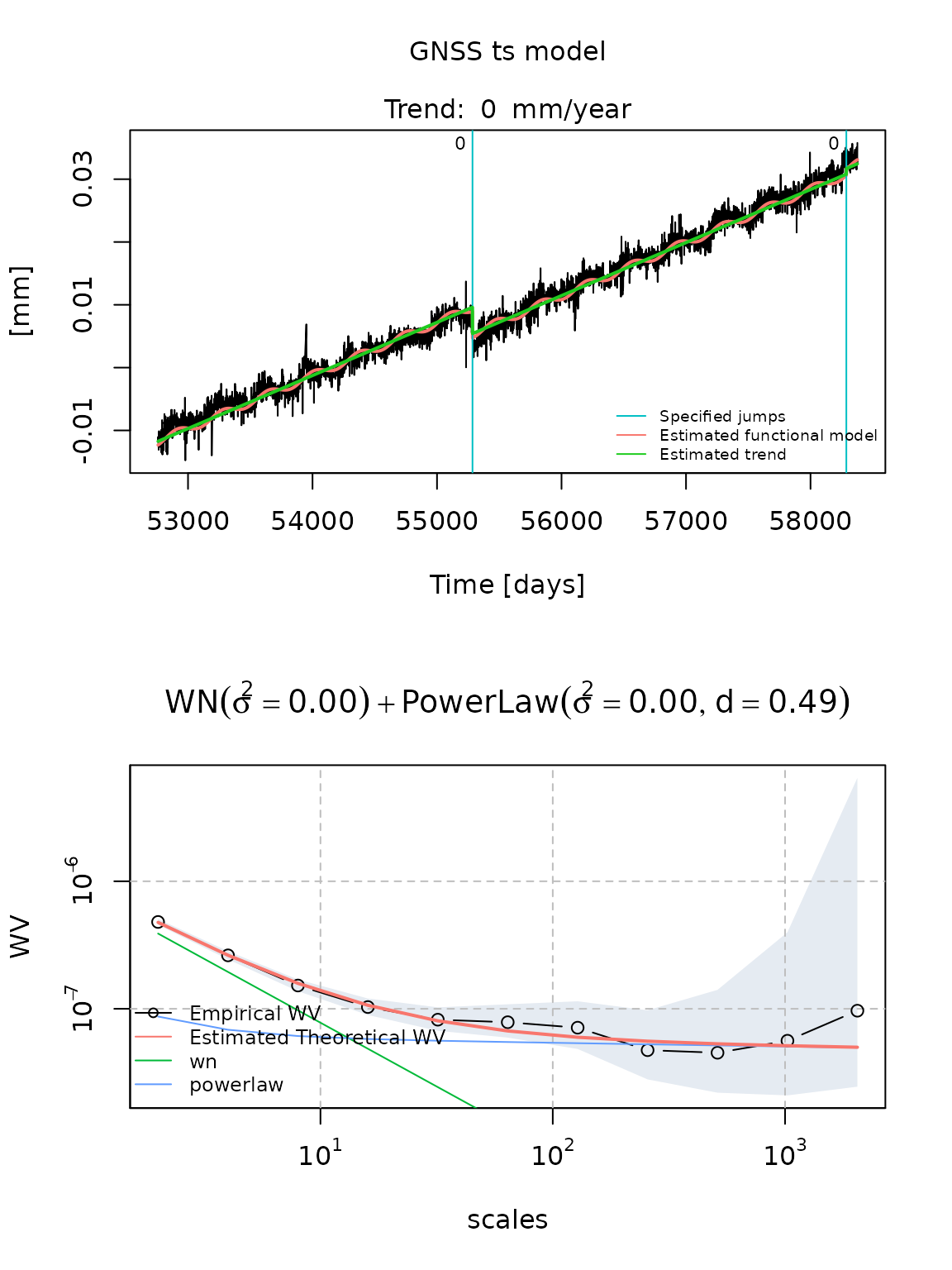
fit_dobs_wn_plp_mle$beta_hat## bias trend A*cos(U) A*sin(U) jump
## 1.200000e-02 8.451006e-06 -6.672690e-04 1.359080e-06 -4.194170e-03
## jump
## 9.071300e-04
fit_dobs_wn_plp_mle$theta_hat## wn_sigma2 powerlaw_sigma2 powerlaw_d
## 7.779727e-07 2.700504e-07 4.854970e-01Importing data from the Plate Boundary Observatory (PBO)
We can load time series data from the Plate
Boundary Observatory (PBO) as gnssts object with
PBO_get_station():
cola = PBO_get_station("COLA", column = "dE")Comparing estimated models
Let us consider three potential models for the stochastic model of this signal. More precisely let us consider:
- a White noise and a PowerLaw process
- a White noise coupled with a Matérn process
- a White noise and a Fractional Gaussian noise
fit_cola_wn_plp = estimate_gmwmx(cola, model_string = "wn+powerlaw",
theta_0 = c(0.1,0.1,0.1),
n_seasonal = 1,
ci = T)
plot(fit_cola_wn_plp)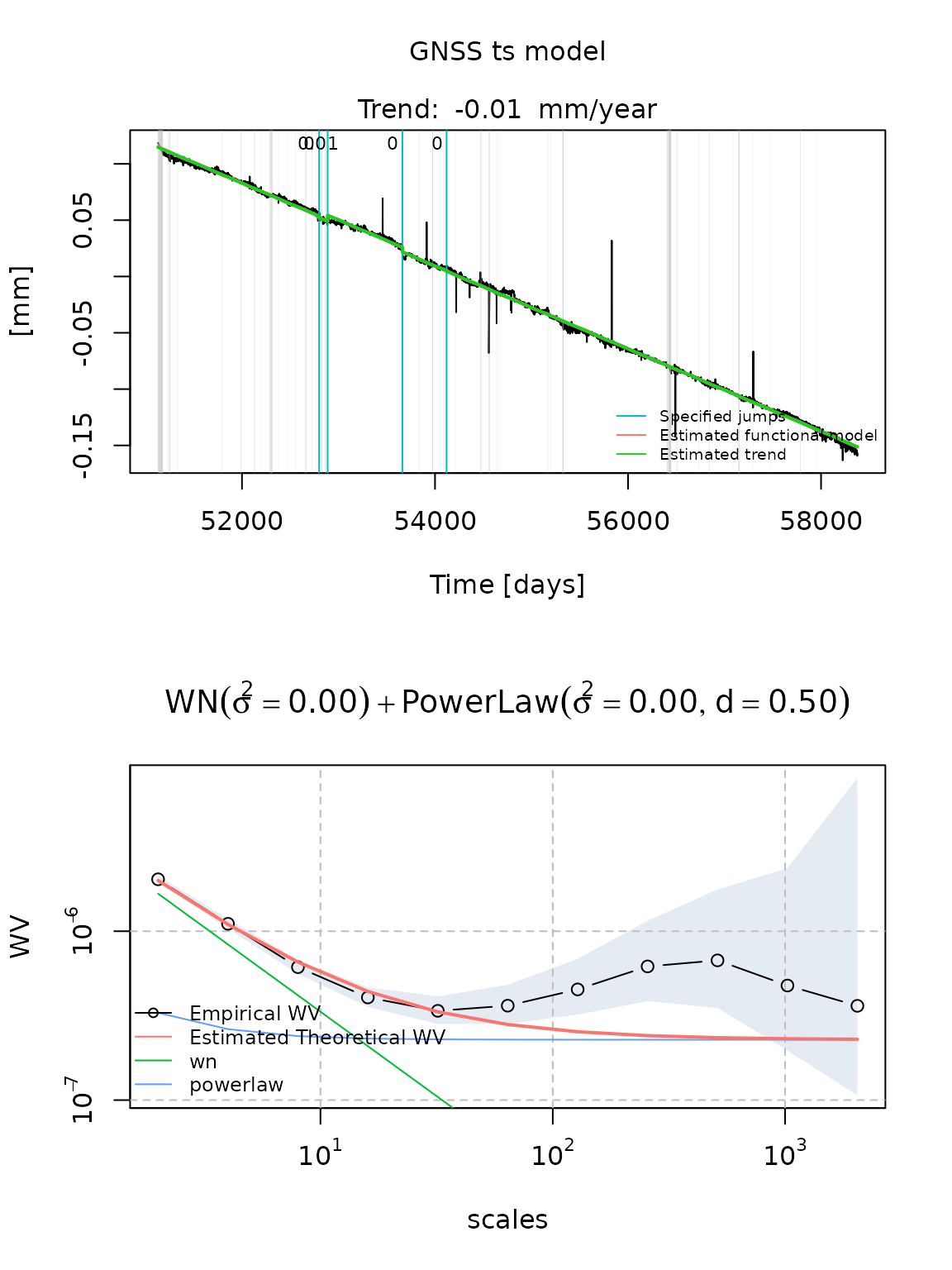
fit_cola_wn_fgn = estimate_gmwmx(cola, model_string = "wn+fgn", theta_0 = c(0.1,0.1,0.2),
n_seasonal = 1,
ci = T)
plot(fit_cola_wn_fgn)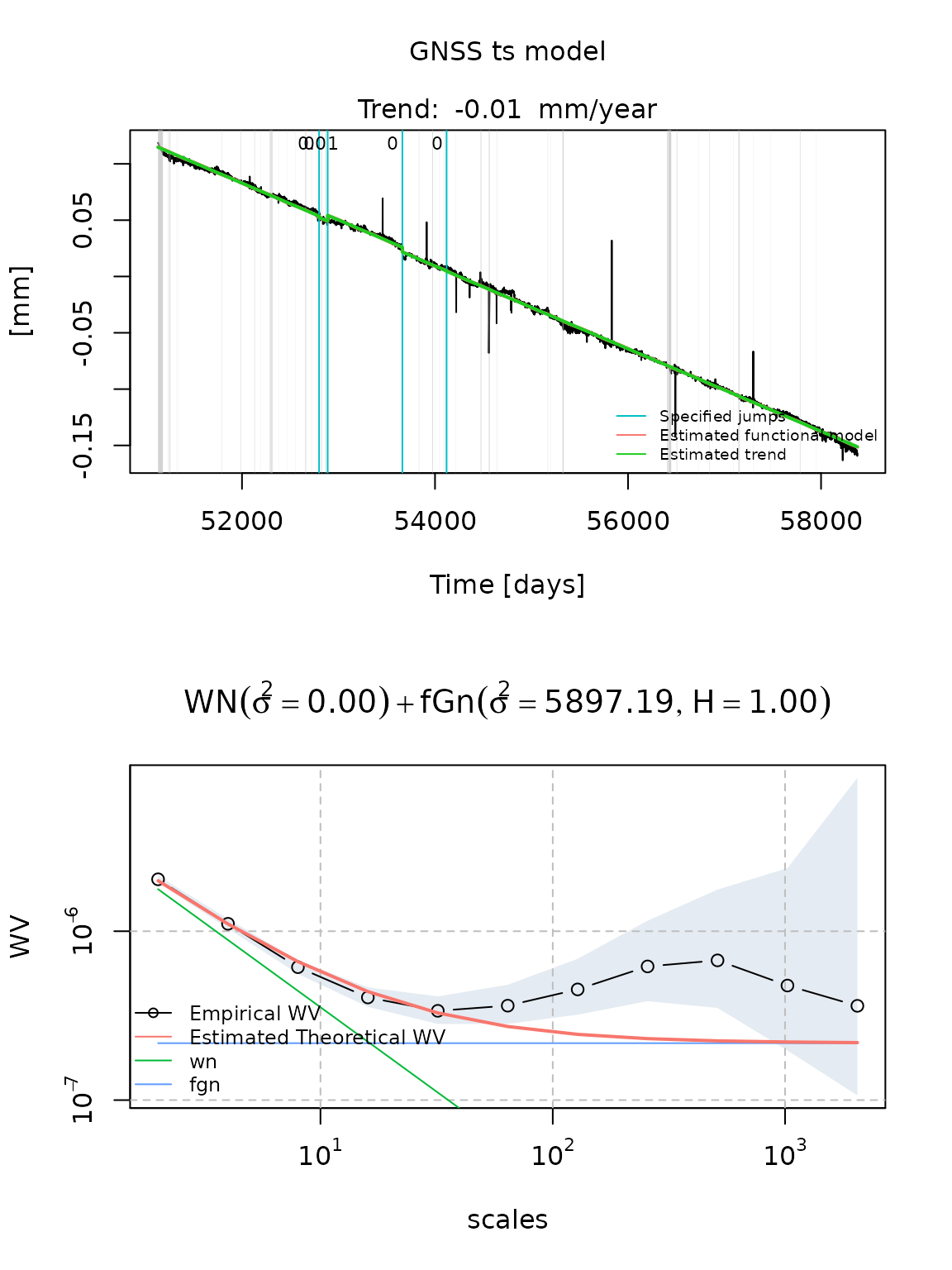
fit_cola_wn_matern = estimate_gmwmx(cola, model_string = "wn+matern",
theta_0 = c(0.1,0.1,0.1,0.1),
n_seasonal = 1,
ci = T)
plot(fit_cola_wn_matern)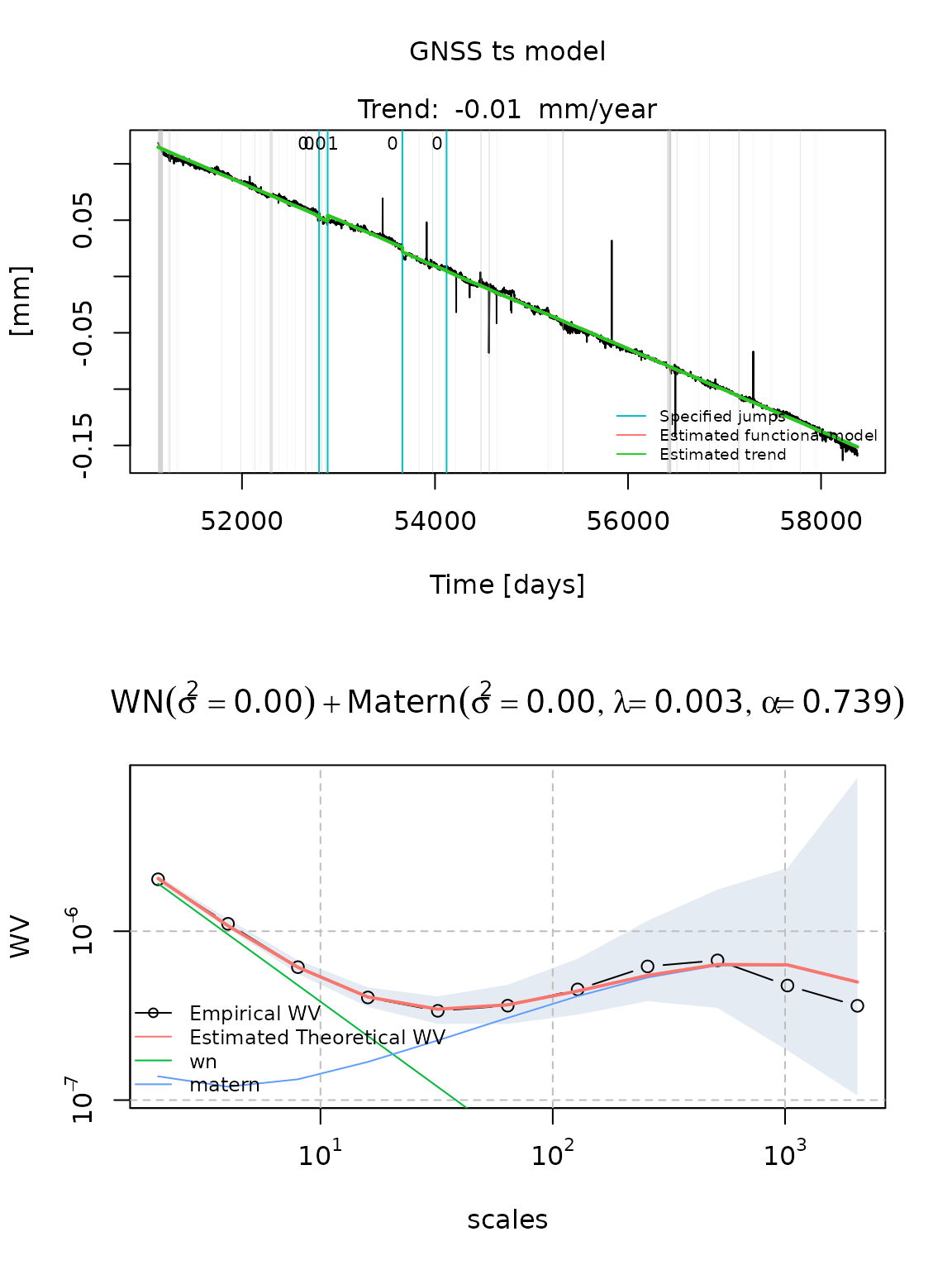
You can compare estimated models with the function
compare_fits()
compare_fits(fit_cola_wn_plp, fit_cola_wn_matern)## Warning in compare_fits(fit_cola_wn_plp, fit_cola_wn_matern): Provided fits do
## not esimate the same model.